Individual Analysis for Personalized Medicine
As scientific and technological advances transform healthcare, individual genomes can be analyzed with the same core technology being used for family medical genetics.
Singleton Accuracy
- Validated true positive detection
- Order-of-magnitude less false positives
- Seamless integration with downstream interpretation
Exomes, Whole Genomes, Panels. It's Your Call.
RTG Singleton uses a Bayesian method (Marth et al. Nat. Genet., 1995) and incorporates priors based on polymorphism rates published by the 1000 Genomes Project. Platform-specific error rates are included and specified in a diploid model, and mapping quality values are also incorporated. Complex regions are identified by various criteria, mainly including regions with apparent indels, MNPs, or clusters of SNVs. A specialized version of our caller for complex regions is used which considers hypothesis including indels and MNPs without the need of realignment. The result is a product that delivers incredibly accurate true variant reporting without letting in significant numbers of false positives that are concomitant with other approaches.
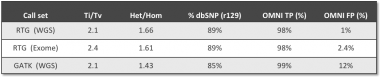
Numbers don't lie. RTG Singleton delivers highly accurate results for targeted panel, exome and whole genome sequence across a range of coverage depths.
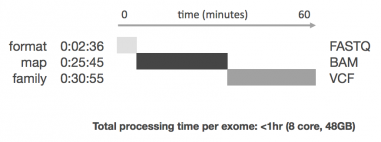
Enterprise Accuracy at the Speed of Now
The RTG Singleton product delivers a 10X improvement on time-to-result for individual sample analysis without requiring special hardware or a massive grid infrastructure. For a single exome at 60X average coverage the product delivers results in under an hour, and the core technology has been fully parallelized so as to leverage as many cores and RAM as are available, meaning multiple individual genomes or exomes can be analyzed using a single infrastructure environment with no tradeoff to per-sample speed.
Local or Cloud. It's your call.
Our philosophy is to move the analytics to your data, not the other way around. While this helps minimize latency and data transfer cost, it also keeps the data where you need it and in compliance with your security requirements. Requiring no hardware purchase, we offer both local "imbedded" pipelines that can be run behind your firewall, and off premises cloud-based pipelines that run in Amazon or can be hosted as a service. You tell us which setup is optimal and we'll take care of the rest.